WolfArray — Spatial Analysis
Note — All operations shown here work identically on
WolfArrayModel(no GUI dependency). See the Model/GUI architecture tutorial for details.
This tutorial covers spatial analysis operations on WolfArray:
Raster algebra (combining arrays with arithmetic operators)
Masking chains (value-based and geometry-based)
Resampling / rebinning with different aggregation operators
Gradient / slope computation
Spatial statistics
[1]:
from wolfhece.wolf_array import WolfArray, header_wolf
import numpy as np
import matplotlib.pyplot as plt
Setup: Synthetic DEM
[10]:
h = header_wolf()
h.shape = (200, 200)
h.set_resolution(1., 1.)
h.set_origin(0., 0.)
dem = WolfArray(srcheader=h)
# Gaussian hill + valley
x = np.linspace(-3, 3, 200)
y = np.linspace(-3, 3, 200)
X, Y = np.meshgrid(x, y, indexing='ij')
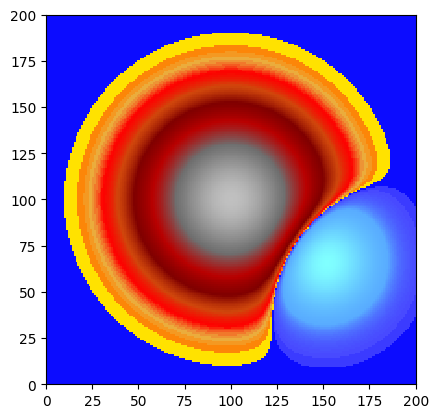
dem.array[:, :] = 100. + 30. * np.exp(-(X**2 + Y**2)) - 10. * np.exp(-((X-1.5)**2 + (Y+1)**2)/0.5)
dem.plot_matplotlib(first_mask_data=False)
[10]:
(<Figure size 640x480 with 1 Axes>, <Axes: >)
Raster Algebra
WolfArray supports full arithmetic: +, -, *, /, **, //, unary -, abs().WolfArray.[11]:
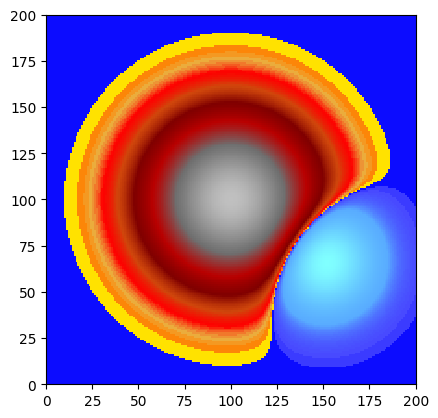
# Compute deviation from mean
mean_val = float(dem.statistics()['Mean'])
deviation = dem - mean_val # scalar subtraction → new WolfArray
deviation.plot_matplotlib(first_mask_data=False)
print(f"Mean of deviation: {deviation.statistics()['Mean']:.6f} (≈0)")
INFO:root:No selection -- statistics on the whole array
INFO:root:No selection -- statistics on the whole array
Mean of deviation: -0.000005 (≈0)
[12]:
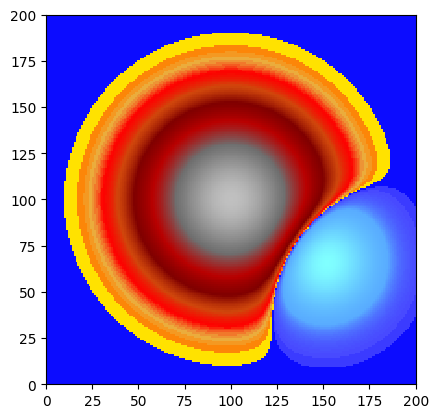
# Normalise to [0, 1]
stats = dem.statistics()
vmin, vmax = float(stats['Min']), float(stats['Max'])
normalised = (dem - vmin) / (vmax - vmin)
print(f"Range: {normalised.statistics()['Min']:.4f} – {normalised.statistics()['Max']:.4f}")
normalised.plot_matplotlib(first_mask_data=False)
INFO:root:No selection -- statistics on the whole array
INFO:root:No selection -- statistics on the whole array
INFO:root:No selection -- statistics on the whole array
Range: 0.0000 – 1.0000
[12]:
(<Figure size 640x480 with 1 Axes>, <Axes: >)
Masking Chains
Multiple mask operations can be chained to isolate specific regions by value.
[13]:
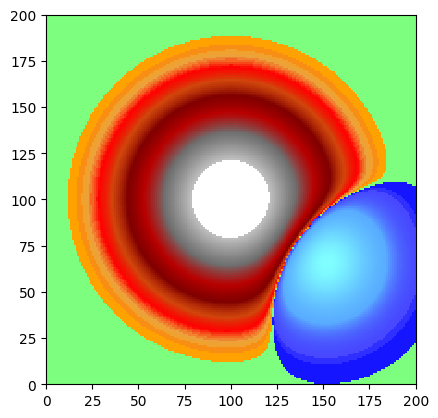
# Isolate elevations between 105 and 120
band = WolfArray(mold=dem)
band.mask_lower(105.) # mask everything below 105
band.mask_greater(120.) # mask everything above 120
band.plot_matplotlib(first_mask_data=False)
print(f"Cells in [105, 120]: {band.count()}")
Cells in [105, 120]: 38622
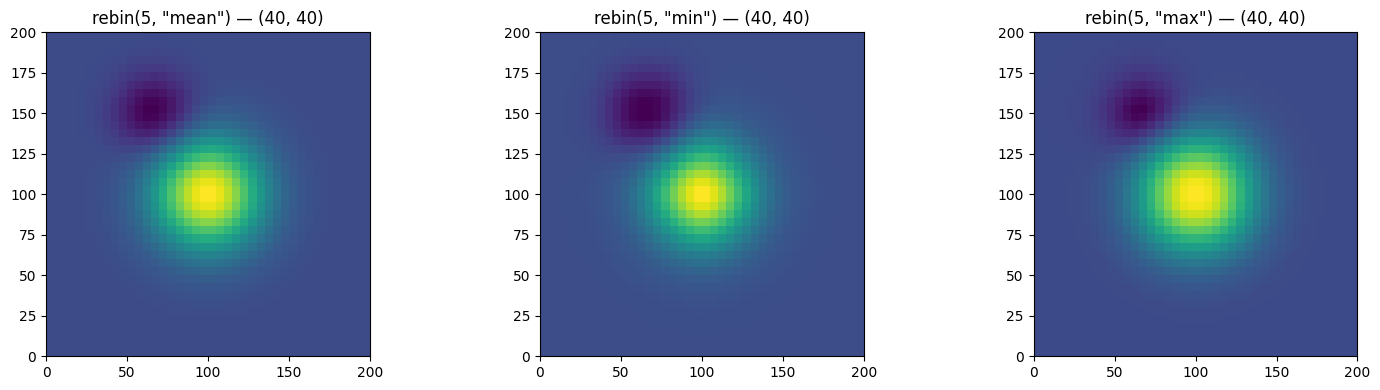
Rebinning with Different Operators
rebin(factor, operation=...) changes the resolution in-place:
factor > 1→ coarser (aggregation)factor < 1→ finer (disaggregation)
Supported operations: 'mean', 'sum', 'min', 'max', 'median'.
[14]:
fig, axes = plt.subplots(1, 3, figsize=(15, 4))
for ax, op in zip(axes, ['mean', 'min', 'max']):
coarse = WolfArray(mold=dem)
coarse.rebin(factor=5, operation=op)
ax.imshow(coarse.array[::-1], extent=[coarse.origx, coarse.origx + coarse.nbx * coarse.dx,
coarse.origy, coarse.origy + coarse.nby * coarse.dy])
ax.set_title(f'rebin(5, "{op}") — {coarse.array.shape}')
plt.tight_layout()
plt.show()
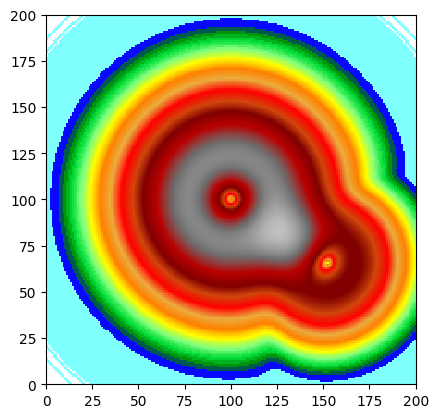
Gradient / Slope
A simple gradient magnitude can be computed using numpy.gradient on the internal array, scaled by the pixel size.
[15]:
# Compute gradient magnitude
gz, gy = np.gradient(dem.array.data, dem.dx, dem.dy)
slope_mag = np.sqrt(gz**2 + gy**2)
slope = WolfArray(mold=dem)
slope.array[:, :] = slope_mag
slope.plot_matplotlib()
print(f"Slope range: {slope.statistics()['Min']:.4f} – {slope.statistics()['Max']:.4f}")
INFO:root:No selection -- statistics on the whole array
INFO:root:No selection -- statistics on the whole array
Slope range: 0.0000 – 0.9372
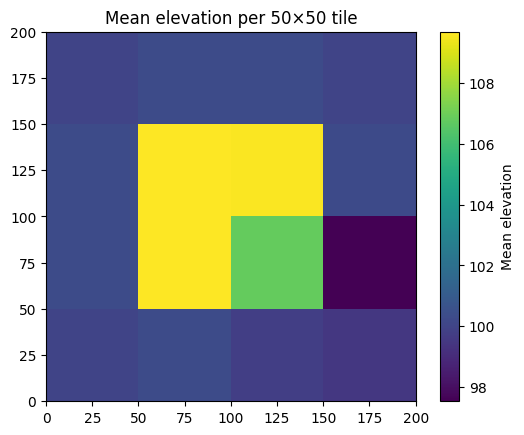
Spatial Statistics by Region
Combining statistics(inside_polygon=...) with systematic spatial partitioning yields a spatial analysis.
[8]:
from shapely.geometry import box
# Divide the domain into a 4×4 grid and compute mean elevation per tile
tile_size = 50
means = np.zeros((4, 4))
for i in range(4):
for j in range(4):
poly = box(i * tile_size, j * tile_size,
(i + 1) * tile_size, (j + 1) * tile_size)
s = dem.statistics(inside_polygon=poly)
means[i, j] = s['Mean']
fig, ax = plt.subplots()
im = ax.imshow(means.T, origin='lower', extent=[0, 200, 0, 200])
plt.colorbar(im, ax=ax, label='Mean elevation')
ax.set_title('Mean elevation per 50×50 tile')
[8]:
Text(0.5, 1.0, 'Mean elevation per 50×50 tile')
Cropping and Edge Trimming
crop_array(bbox)— extract a subregion by world coordinatescrop_masked_at_edges()— automatically trim rows/columns that are fully masked at the edges
[9]:
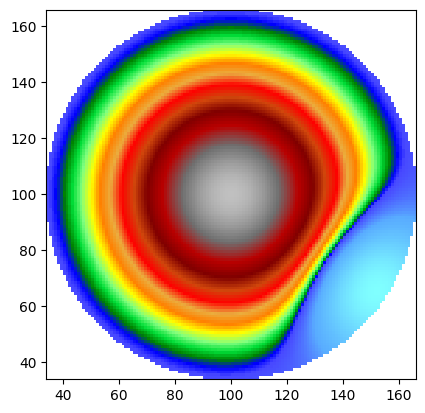
# Mask outside a circle, then trim
circle_dem = WolfArray(mold=dem)
dist = np.sqrt((X - 0.)**2 + (Y - 0.)**2)
circle_dem.array[dist > 2.] = circle_dem.nullvalue
circle_dem.mask_data(circle_dem.nullvalue)
trimmed = circle_dem.crop_masked_at_edges()
print(f"Before trim: {circle_dem.array.shape}")
print(f"After trim: {trimmed.array.shape}")
trimmed.plot_matplotlib()
Before trim: (200, 200)
After trim: (132, 132)
[9]:
(<Figure size 640x480 with 1 Axes>, <Axes: >)